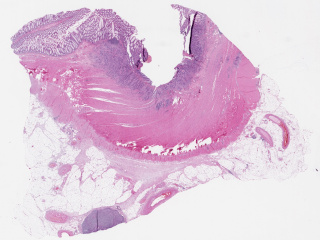
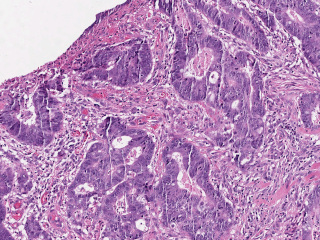
Whole slide pathology images from regional lymph node metastasis in colon adenocarcinoma produced at Region Gävleborg Clinical Pathology and Cytology department and Region Östergötland Clinical Pathology department. Annotations for AI training produced as part of AIDA clinical fellowship project investigating AI decision support in metastasis detection.
This dataset has been extended with a second collection series in the LNCO2 dataset using different collection and annotation parameters.
Keywords: Pathology, Whole slide imaging, Annotated, Lymph nodes, Cancer, Colon, Adenocarcinoma.
Sample images
Sample images with reduced image quality. Please click to preview.
Dataset information
| Short name | LNCO |
|---|---|
| Origin | Clinical |
| Cite as |
Gordan Maras, Martin Lindvall, and Claes Lundstrom
(2019)
Regional lymph node metastasis in colon adenocarcinoma
doi:10.23698/aida/lnco [BibTeX format] |
| Field | Pathology |
| Organ |
Colon |
| Age span | |
| Title | Regional lymph node metastasis in colon adenocarcinoma |
| Author |
Gordan Maras
Martin Lindvall Claes Lundstrom |
| Year | 2019 |
| DOI | doi:10.23698/aida/lnco |
| Status | Completed |
| Version | 1.2.0 |
| Scans | 977 |
| Annotations | 7379 |
| Size | 373.66GB |
| Resolution | 20x and 40x |
| Modality |
SM
|
| Scanner |
Aperio Scanscope (20x) Hamamatsu NanoZoomer (40x) |
| Stain | Hematoxylin and eosin. |
| Phase | |
| References |
|
| Copyright | Copyright 2019 Linköping University, Claes Lundström |
| Access |
Available under the following licenses, described in the License section below.
Controlled access
AIDA BY license
|
Annotation
ROI boxes around lymph nodes (positive, negative or unasssed) and detailed tumor polygons
See details below.
Annotation details
Labels
The following labels exists:
-
roi_lgl_norm:: Region contains a lymph node section that do not contain tumor-positive areas, eg, is normal or negative. (but see also note below)
-
roi_lgl_unknown:: Region contains a lymph node for which ground truth has not been established.
-
roi_lgl_tumor:: Region contains a lymph node where at least some areas are tumor-positive areas. (Might not be annotated)
-
roi_lgl_exclude:: Contains lymph node that was determined to be of poor quality or contain severe artifacts from scanning. Would be excluded from clinial review (not suitable).
-
tumor:: region exclusively contains tumor pixels (but see also excl category below)
-
excl & excl_artifact:: these areas should be substracted from tumor-annotated regions to gain pure-tumor pixels.
-
bkg: region contains only background pixels
NOTE: tumor annotations are not exhaustive, meaning, there might be more tumor areas in lgl_roi_tumor lymph nodes than annotated with tumor annotations.
NOTE: Some roi_lgl_norm areas contain tumor cells that are from an accidental contamination of the glass during processing. This is normal during routine clinical work and a challenge that automated systems should deal with in an appropriate manner. This is a very easy task for human pathologists to discern.
NOTE: tumor regions not within a roi_lgl_* region should be assumed to be annotations of tumor in the the primary tumor. This can easily be verified visually, and the primary tumor is always on the initial blocks of the case (A..B…C etc)
Sampling procedure
Sampling procedure: Query for patients with confirmed adenocarcinomas that underwent colorectomy with subsequent lymphnode review as part of TNM-staging. Query selected N cases with a relatively high number of reported positive lymph nodes.
Source: Two medical clinics in Sweden.
Descriptive statistics
LNCO1
patients: 39
cases: 39
images: 977
median dimensions(WxH px): 49799 x 38729
median resolution (micrometer per pixel): 0.4959
annotations:
number: 7379
lymph node sections (roi) 1557
which are negative 712
which are positive 292
which are unassessed 525
images with at least one detailed tumor annotation: 165
File formats
Pixel position scaling
Coordinates given are relative to the image width. To get the correct pixel position, X coordinates (and Y coordinates!) should therefore be multiplied with the image width.
License
Controlled access
Free for use in legal and ethical medical diagnostics research.
To request access to the dataset, use the Apply for access button below.
Note that the recipient researcher must hold at least a PhD degree in a
relevant field and that the applicant should be an authorized signatory who can
legally enter into data sharing agreements on the behalf of the institution.
Help with applying for controlled access.
AIDA BY license
Copyright 2019 Linköping University, Claes Lundström
Permission to use, copy, modify, and/or distribute this data within Analytic Imaging Diagnostics Arena (AIDA) for the purpose of medical diagnostics research with or without fee is hereby granted, provided that the above copyright notice and this permission notice appear in all copies, and that publications resulting from the use of this data cite the following works:
Gordan Maras, Martin Lindvall, and Claes Lundstrom (2019) Regional lymph node metastasis in colon adenocarcinoma doi:10.23698/aida/lnco.
THE DATA IS PROVIDED “AS IS” AND THE AUTHOR DISCLAIMS ALL WARRANTIES WITH REGARD TO THIS DATA INCLUDING ALL IMPLIED WARRANTIES OF MERCHANTABILITY AND FITNESS. IN NO EVENT SHALL THE AUTHOR BE LIABLE FOR ANY SPECIAL, DIRECT, INDIRECT, OR CONSEQUENTIAL DAMAGES OR ANY DAMAGES WHATSOEVER RESULTING FROM LOSS OF USE, DATA OR PROFITS, WHETHER IN AN ACTION OF CONTRACT, NEGLIGENCE OR OTHER TORTIOUS ACTION, ARISING OUT OF OR IN CONNECTION WITH THE USE OR CHARACTERISTICS OF THIS DATA.